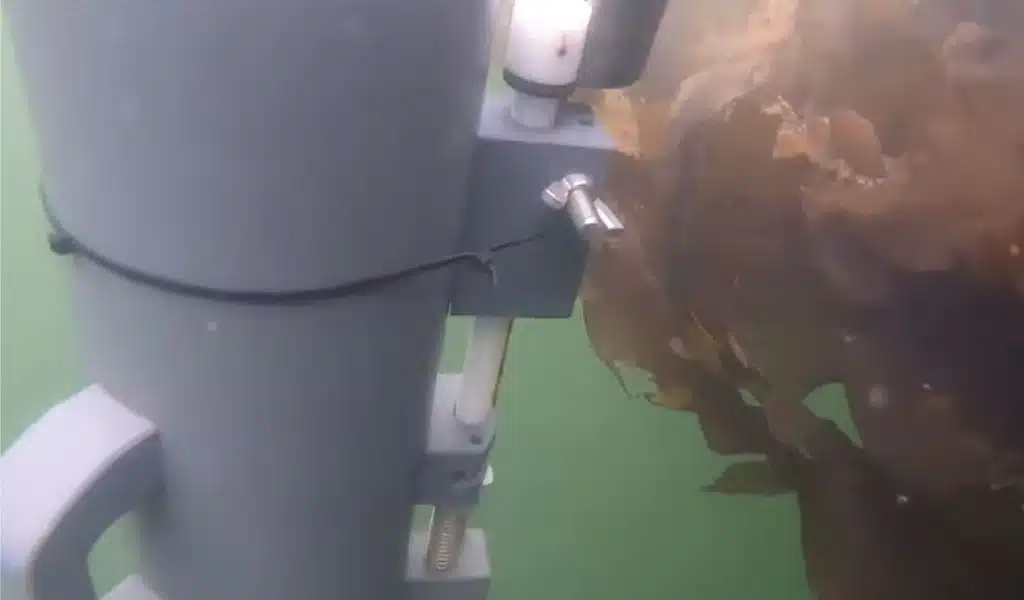
Nahi El Bar Jiyed hangs off the side of the boat and fills a jug with sea water. He then carefully pours the water into a big syringe. Even though the sample doesn’t look like much, it contains the DNA of every living thing in the water below.
He puts a sample of water through a screen about the size of his hand, which catches the DNA pieces. He does this process several more times. A sea turtle comes out of the water to take a breath and then goes back down to the seagrass patch below.
Jiyed says, “We can take a sample that will tell us exactly what was at this site without disturbing the environment.”
In the last ten years, the use of environmental DNA, also called “eDNA,” to track biodiversity has grown a lot. As animals move through their surroundings, they lose pieces of their DNA, such as skin cells, waste, and other body fluids. By getting these tiny bits of DNA from samples of water, dirt, or air, scientists can find out with an accuracy never before seen how many species are there and how different they are. This gives a snapshot of how complicated an ecosystem is.
“Management is based on what you know. Jiyed says, “If you don’t know a place, you can’t protect it.” “It’s just the first step.” His work is part of a Unesco project to collect eDNA from 22 marine world heritage places, such as the national park where he works in Mauritania, Banc d’Arguin.
The park is on the north side of the country, where Saharan sand meets green waters. Low-lying islands are passed by fishing boats that move only by the wind. The deep quiet is deceiving. This place is home to dolphins and sea turtles that are in danger of extinction, and it is an important stopover for millions of migrating birds. In 1989, Unesco named the park a world heritage site because it had so many different kinds of plants and animals.
The park head, Ebaye Sidina, calls it a “true biogeographic crossroads” where tropical and temperate organisms meet. “We know there are a lot of species, and the DNA analysis will finally pull back the curtain and show that this diversity exists and needs to be protected,” he says.
Scientists can use eDNA technology to measure biodiversity, find invasive species, track rare or hard-to-find animals, and look for diseases and pathogens in wastewater. It has even shown that some species that were thought to be dead are still alive. Scientists are looking for signs of the smalltooth sawfish at Banc d’Arguin, which Sidina said hasn’t been seen in decades.
Using eDNA over time can show how the climate problem is affecting populations, such as by changing where they live.
“Several fish species are already moving 25 km every 10 years, either to deeper waters or farther from the equator,” says Fanny Douvere, head of Unesco’s world heritage marine program. “We want to make sure that these marine world heritage sites will still have the same boundaries in 30 to 40 years.”
Analysis of eDNA is better than standard surveys in a number of ways, especially when figuring out how climate change affects an ecosystem. Ward Appeltans, head of the ocean biodiversity information system at Unesco and science coordinator of the eDNA expeditions project, says that eDNA analysis can give results in just a few months, while traditional surveys can take years to finish.
The condition of the water is changing so quickly, he claims. We want to know the situation today, not five years ago.
Tracking invasive species, like the lionfish, which can quickly displace local fish populations, requires the capacity to recognize sudden changes. Additionally, non-invasive eDNA sampling provides an alternative to hazardous conventional survey techniques like bottom trawling and the capture, tranquilization, and tagging of animals.
According to Luke Thompson, a researcher at Mississippi State University and the National Oceanic and Atmospheric Administration in Miami, Florida, “eDNA will also pick up things that you maybe wouldn’t have seen because they were hiding or only show up at certain times of the day.” Traditional marine and airborne surveys are typically far more expensive as well because they need boats, helicopters, planes, and staff.
But eDNA technology does have some problems of its own. Ocean currents can make it hard to find species in the sample area or make species from far away look like they are there. Some animals lose more DNA than others, which can make it hard to figure out how many of each animal there are.
But, says Louis Bernatchez, editor-in-chief of the scientific magazine Environmental DNA, the biggest problem is that there is no agreement on how to standardize methods and markers.
Scientists use a reference database to match the genetic patterns of the DNA they have found to the species they belong to. But there isn’t a single world database, and there’s no agreement on which genetic markers to use. As of now, the genes of less than 1% of all animals have been sequenced.
Bernatchez says, “People aren’t working together enough.” “It’s a very big problem.” Unesco, on the other hand, wants to add all of the information from its eDNA project to its open science marine species library.
Even though there are problems, experts all over the world continue to find new ways to use the technology. In the Arctic, DNA from polar bear tracks is being used to keep track of population size and movement patterns. This knowledge could help conservation efforts and make it easier for people and bears to live together.
At the bottom of the ocean is a purple sea creature called a “gummy squirrel.” It is see-through.
In a deep-sea mining hotspot in the Pacific, more than 5,000 new species have been found.
Read on…
eDNA samples from flowers in North America and Europe have found animals and insects that were not known to be pollinators before. In the field of forensics, scientists have found that taking DNA from the blood of mosquitoes at crime scenes can properly identify victims and suspects.
Thompson says, “There are a lot of different molecules out there that are just waiting to tell us what’s going on and what they’re doing.” “That’s an excellent way to look at it.”